A comparison of Mass Spectroscopy and High Performance Capillary Electrophoresis (HPCE) in the quality control analysis of glycoproteins.
Introduction:
Ribonuclease B glycoform separations were used as benchmark molecule in a comparative study of analytical techniques used in glycosylation profiling. Techniques compared included deltaDOT’s HPCE system and Mass Spectrometers. The data presented here compares deltaDOT’s HPCE with the Agilent 6530 Q-TOF system.
Challenge:
The next 10-12 years are set to see a dramatic increase in the use of biological drugs with monoclonal antibodies and their associated biosimilars/betters leading the way. In addition, currently available biological drugs require complex process analytics at many stages during their production. These stages include up- mid- and downstream, QC/QA through to formulation and batch release. Within each stage varying analytical needs include measurement of: molecular weight, concentration, fragmentation, primary, secondary and tertiary structure and any relevant post-translational modifications. These analytical requirements generate high demands and associated costs for manufacturers. With the UK government’s expressed wish to increase the level of stratified medicines over the next 15 years, and the ideal of near to patient manufacture, the world of bioproduction analytics has extensive challenges to overcome.
Rationale:
For many years Mass Spectrometry has been correctly regarded as the gold standard in separation technology. Its mass assignment can be to 4 or five decimal places and the existing databases allow seamless identification of analytes. It is not however, without its associated problems, such as capital expenditure, operational expenditure such as servicing and software packages and experienced personnel, – critical in data interpretation. Availability of Mass Spectrometry instruments even in the largest biopharmaceutical companies can be a bottleneck, with waiting times typically of two weeks for samples to be run. Other techniques providing data of adequate analytical quality are scarce. PAGE-gels cannot easily or routinely produce the required resolution neither can HPLC. Canonical capillary electrophoresis has issues with user experience, repeatability and resolution at this level of analysis.
deltaDOT solution:
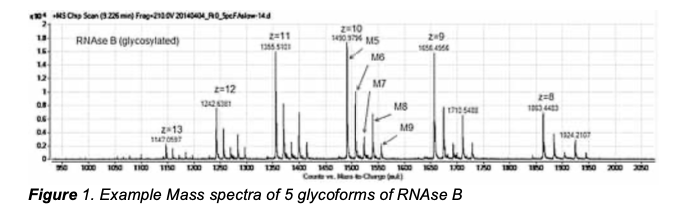
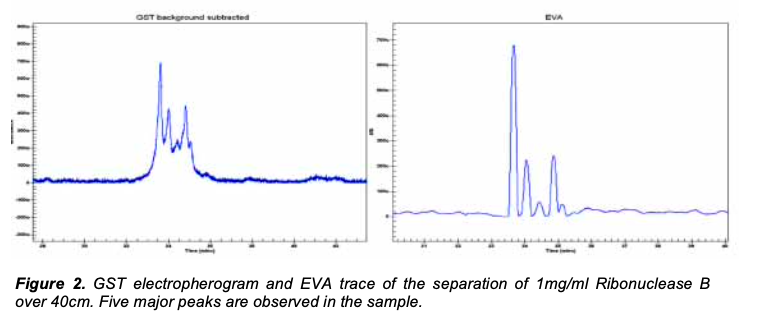
deltaDOT’s technology is significantly different to conventional Capillary Electrophoresis (CE) in which the separated components are detected by a single point detector after separation. It is through gathering data from 512 points, and using proprietary signal processing algorithms, that deltaDOT’s HPCE is able to produce higher quality results than competing CE systems. Ribonuclease B (SIGMA) was analysed on an Agilent 6530 QTOF* with chip cube technology, Agilent Zorbax 300SB-C18 chip using a 5µm 30Å 75µm x150mm with 40nL enrichment column (Figure 1) and deltaDOT’s HPCE system with 512 diode array detector 50cm, 50µm bare fused silica capillary in an SDS run buffer (Figure 2).
Analysis of the HPCE separations was performed using deltaDOT’s Equiphase Vertexing Algorithm (EVA) and Generalised Separation Transform (GST) algorithm. GST is a method of combining the data from the 512 pixels which preserves the peak shape while at the same time maximising the signal-to-noise ratio. EVA is a technique developed by deltaDOT which uses a pixel-time constraint to improve the signal-to-noise which improves the resolving power of the system The results quoted in this report are based upon the GST electropherograms and EVA-analysed traces.
The data in Figure 1 produced by the Q-TOF is complex owing to the presence of multiply charged ions. For each charge, the relative heights of the peaks change, thus quantitation of each varies depending on the charge. In comparison, the data in Figure 2 using the EVA algorithm is able to resolve the peaks with baseline separation. It is also able to quantify each glycoform with excellent repeatability.
The independent Mass Spectrometry data shown in Figure 1 was produced at the Laboratory of the Government Chemist (LGC). A poster comparing the data from both techniques concluded that the data produced by the Agilent 6530 QTOF and deltaDOT’s HPCE system on the Ribonuclease B Glycoforms were comparable. This is a significant outcome as Agilent 6530 QTOF costs c. 3-4 times more to buy and has significantly higher running costs compared to deltaDOT’s HPCE.
Commercial applications:
The high quality data produced by deltaDOT’s HPCE can been successfully used as an alternative to Mass Spectrometry in many areas, as follows.
- Comparison of bioreactor growth parameters for glycosylated therapeutics.
- Allowing the analysis of glycosylated therapeutics in a timely fashion throughout the production process.
- Analysis of stability in glycosylated therapeutics at point of delivery.
- Allowing distributed analysis of lifestyle markers.
- Providing a “triage” on sample quality prior to expensive orthogonal methods
- Reducing pressure on over-worked Mass Spectrometers
Further information
For any additional information on our ability to characterise complex samples please contact us on info@deltadot.com. Click here for referenced poster produced by LGC.
*Acknowledgements: Sabine Biesenbruch, Milena Quaglia, Kate Groves, Frank Torma, LGC, Queens Road, TW11 0LY, Teddington, UK
