Antibiotic analysis Introduction: Analysis of ß-lactam antibiotics in food products can be performed using the deltaDOT HPCE. This enables the possibility of on-site analytics of at food at source or at distribution centres. Various penicillin derivatives are widely used to treat infectious diseases in humans and animals, acting to kill or inhibit the growth of the infectious organism. A mixture of these 8 antibiotic derivatives was analysed using Micellar Electrokinetic Capillary Chromatography.
October 2017 deltaDOT LONDON
Challenge: The presence of antibiotics in the food chain is tightly controlled to avoid new strains of antibiotic resistant bacteria developing. EU regulations now determine maximum limits for each of the 8 penicillin derivatives that can be allowed in animal tissue products for the consumer. What is needed is a simple, repeatable and cost efficient way to analyse the presence and identity of antibiotics in the food chain from farm to sale point. As with many chemical entities or small molecules, the analysis of antibiotics is largely dependent on established techniques such as ELISA (an antibody based kit), High Performance Liquid Chromatography and Mass Spectrometry. While all of these techniques have merit, such as pedigree and the associated market acceptance that this brings, they also have problems in expense, skill-set availability and centralization. These lead inevitably to delay and cost implications that are not easily addressed.
Rationale: While antibiotics have been a life-saving therapy since the 1940’s they are not without their problems. Foremost of these is the resistance issue. While it is normal to view bacteria as a single population, in fact this is not the case. A single colony of e.g. E.coli consists of millions of individual bacteria, all constantly growing. Bacteria do have a sexual aspect to their life, as they do not simply divide, but also transmit their DNA into their neighbours. This they do as a matter of course, by injection small packages of DNA, called plasmids, into other bacteria via a spiky projection from their cell wall called a sex pili. Sometime this plasmid is able to integrate into the host’s DNA and change the proteins the bacteria makes, but often not. Given that E.coli replicates every 20 minutes or so, and there are 10s of millions in each colony, it is possible to see that the colony can contain many different variants of the same basic bacteria. A few of these may survive and thrive; the majority fail or show no external difference to the main bacteria population. However, if a stress is put on the colony, such as the presence of an antibiotic, then some of these “new” variants may find themselves in an advantageous position. All these variant bacteria are fighting for the same resources (sugar, oxygen etc.) and if there is a sudden cull of bacteria that are vulnerable to the antibiotic, those that are by chance resistant prosper and grow as they have more resistance. This is why medical doctors often have to try more than one antibiotic on a patient. The constant misuse of antibiotics is such a problem tha prescribing them e.g. for a cold, which is a virus and untouched by antibiotics, means that the cold is untouched. Any bacteria susceptible to that drug is eliminated, however, freeing up food for resistant bacteria. Those bacteria are now what is called drug-resistant and can cause terrible problems. In the human gut, there are huge populations of bacteria, many of which are benign and indeed vital to our metabolism. Some however can be malign under certain circumstances, such as E.coli species, and so the presence of antibiotics in food such as milk must be monitored. This prevents the opportunistic growth of dangerous bacteria and their possible spread into the population. 2 deltaDOT Solution: Micellar Electrokinetic Capillary Chromatography (MECC) High Performance Capillary Electrophoresis. MECC relies on the different mobility of the antibiotics through an environment of very small detergent “bubbles” or droplets. A mixture of 8 Penicillin derivatives was analysed using deltaDOT’s proprietary label-free technology with Multipixel detection (Figure 1). Capillary electrophoresis was used for the separation of (1) Amoxicillin, (2) Ampicillin, (3) Penicillin G, (4) Oxacillin, (5) Penicillin V, (6) Cloxacillin, (7) Nafcillin and (8) Dicloxacillin, using a sodium tetraborate buffer supplemented with sodium dodecyl sulphate (SDS). Sodium dodecyl sulphate (SDS) is the most widely used surfactant in MECC, forming negatively charged droplets. Using constant separation conditions, this allows resolution of this set of weakly acidic compounds based upon their hydrophobicity and their variable partitioning into the hydrophobic core of the droplet. Analysis was performed at pH 8.0, which improves tailing of the peaks and allows the generation of a strong overall electro-osmotic flow towards the anode, in order to allow elution of the hydrophobic compounds while they reside in the aqueous phase.
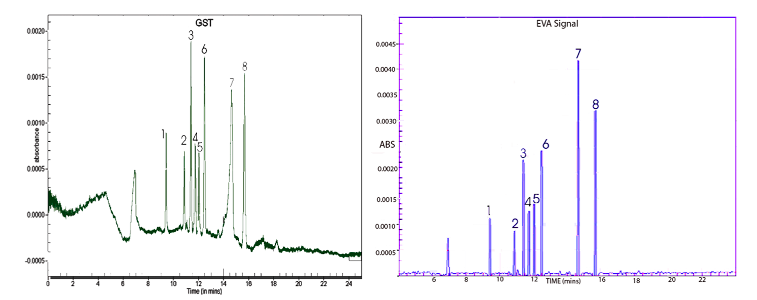
Figure 1. Standard GST electropherogram and EVA trace for the separation of 8 penicillin derivatives. The analytes are (1) Amoxicillin, (2) Ampicillin, (3) Penicillin G, (4) Oxacillin, (5) Penicillin V, (6) Cloxacillin, (7) Nafcillin, (8) Dicloxacillin. For peak 1 the mobility (<0.5% RSD) and peak area (<3.0% RSD) data shows the high quality of the data.
Commercial Applications: The high quality data produced by deltaDOT’s HPCE can be successfully used as an alternative existing, expensive analysis techniques in many areas, as follows: –
• Milk contamination analysis at the distribution or packaging centre
• Detection of antibiotics in milk formula post-formulation
• Detection of antibiotics above the recommended concentration in meat products
• Detection of antibiotics in recycled animal products in animal feed
• Detection of antibody residues in containers used to ship animal food or components e.g. amino acids from third party suppliers
Further information For any additional information on our ability to characterise complex samples please contact us on info@deltadot.com.
